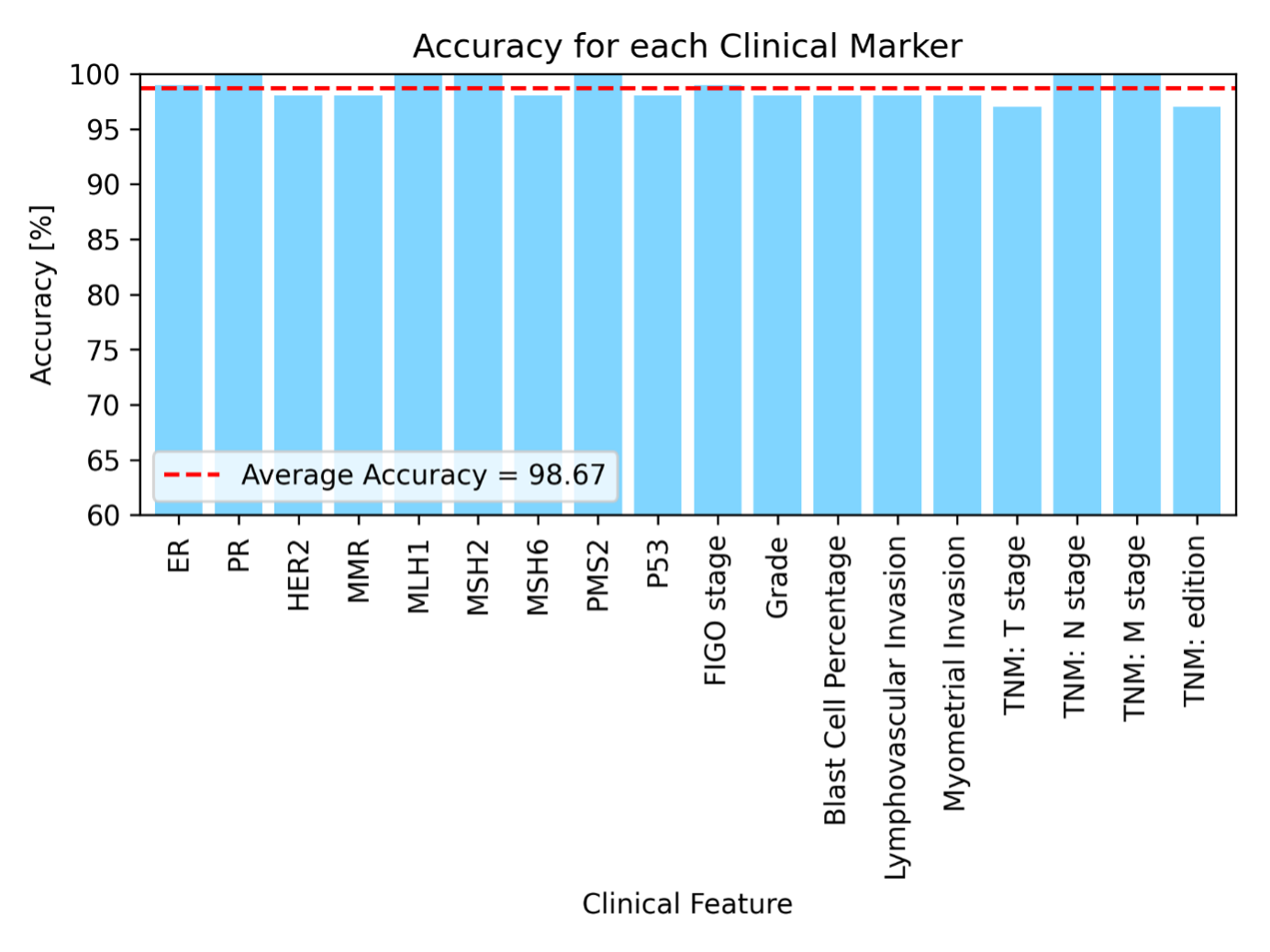
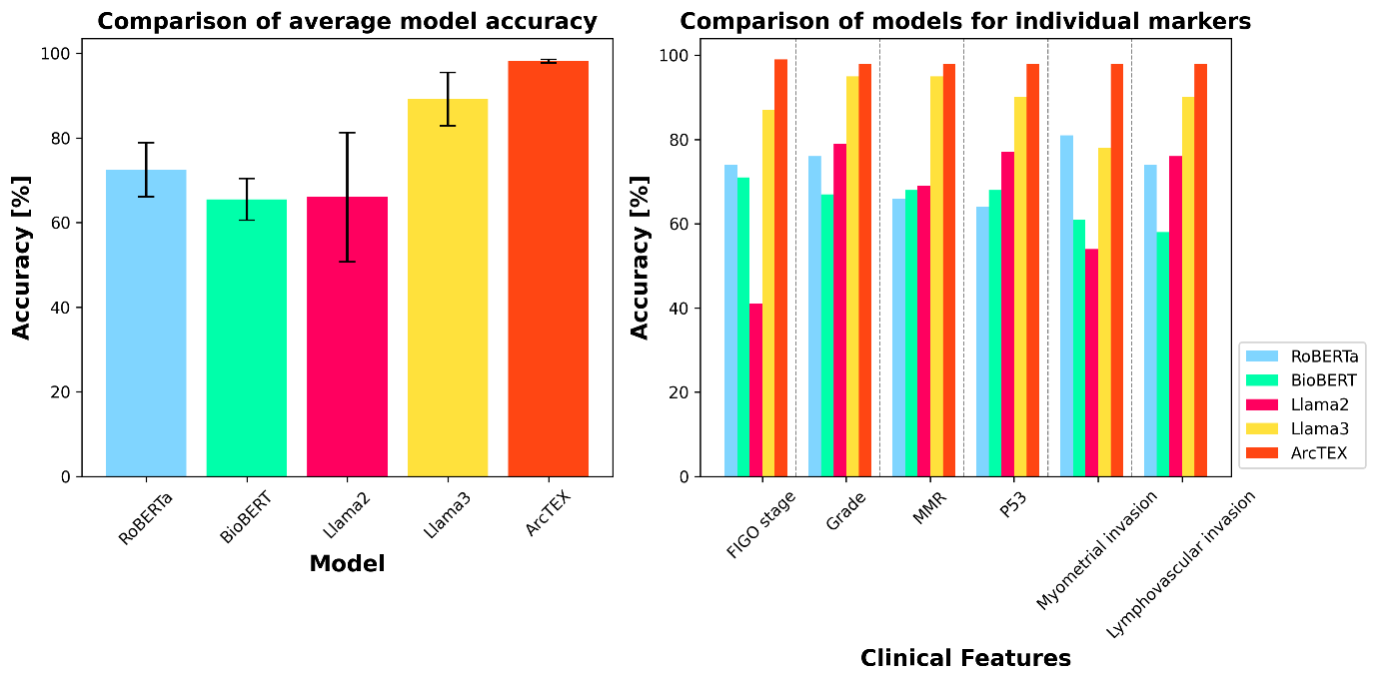
Mismatch Repair (MMR) proteins—MLH1, MSH2, MSH6, and PMS2—play a crucial role in correcting DNA replication errors. Mutations in these proteins can lead to MMR deficiency (dMMR), contributing to genomic instability and impacting treatment decisions in cancers such as endometrial and colorectal cancer. However, identifying patients with dMMR from real-world data is challenging due to the variability in how MMR and its four sub-markers are documented in free-text pathology reports. Reporting practices vary significantly across pathologists, hospitals, and healthcare systems. Some reports provide detailed information on each individual protein, others simply state the overall MMR status as deficient (dMMR) or proficient (pMMR), and some use a combination of these approaches.
Naïve methods, such as using basic search techniques like regular expressions, often struggle to effectively capture this variability, resulting in a limited number of patients being correctly identified for analysis.
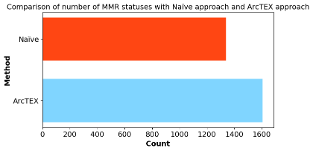
In a recent analysis of an oncology dataset contained within the Arcturis real world data. ArcTEX was able to identify 20% more reports with actionable MMR status compared to a naïve implementation by considering both the overall MMR status and all four submarkers. As shown below, ArcTEX was able to identify 1,606 reports containing MMR information, compared to 1,336 reports using a naïve approach. Additionally, ArcTEX can filter out reports where clinical features are mentioned in a non-actionable context, such as “not applicable,” “requested,” or “to follow.”
Of these 1,606 reports, 51.9% contained both the overall MMR status and details about individual submarkers, 31.4% mentioned only the overall MMR status, and 16.6% listed the individual submarkers without specifying the overall status. This variability highlights the complexities of MMR reporting, even within a single dataset.
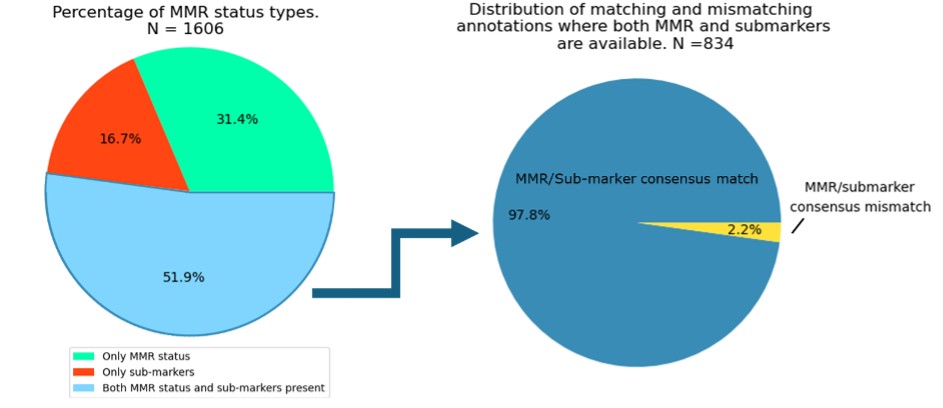
For reports that include both, the overall MMR status and individual submarkers, ArcTEX further enhances data quality by cross validating this information. As illustrated in the figure above, there is a 97.8% agreement between explicitly stated MMR status and the status inferred from the individual submarkers when both are available. The remaining 2.2% of discrepancies are flagged for manual review. This process allows for a more efficient and accurate analysis of free-text oncology data, ultimately enhancing data reliability and providing deeper insights into oncology research.